5. [DocU] Tutorials on using the ADAO module in SALOME¶
This section presents some examples on using the ADAO module in SALOME. The first one shows how to build a very simple data assimilation case defining explicitly all the required input data through the EFICAS graphical user interface (GUI). The second one shows, on the same case, how to define input data using external sources through scripts. We describe here always Python scripts because they can be directly inserted in YACS script nodes, but external files can use other languages.
These examples are intentionally described in the same way than for the [DocU] Tutorials on using the ADAO module in Python because they are similar to the ones that can be treated in the textual user interface in Python (TUI). The mathematical notations used afterward are explained in the section [DocT] A brief introduction to Data Assimilation and Optimization.
5.1. Building an estimation case with explicit data definition¶
This very simple example is a demonstration one, and describes how to set a BLUE estimation framework in order to get the fully weighted least square estimated state of a system from an observation of the state and from an a priori knowledge (or background) of this state. In other words, we look for the weighted middle between the observation and the background vectors. All the numerical values of this example are arbitrary.
5.1.1. Experimental setup¶
We choose to operate in a 3-dimensional observation space, that is we deal with
3 simple measures. The 3 dimensionality is chosen in order to restrict the size
of numerical object to be explicitly entered by the user, but the problem is
not dependent of the dimension and can be set in observation dimension of 10,
100, 1000… The observation  is of value 1 in each
direction, so:
is of value 1 in each
direction, so:
Yo = [1 1 1]
The background state  , which represent some a priori
knowledge or a mathematical regularization, is chosen of value of 0 in each
case, which leads to:
, which represent some a priori
knowledge or a mathematical regularization, is chosen of value of 0 in each
case, which leads to:
Xb = [0 0 0]
Data assimilation requires information on errors covariances 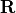 and
and  , respectively for observation and background error
variables. We choose here to have uncorrelated errors (that is, diagonal
matrices) and to have the same variance of 1 for all variables (that is,
identity matrices). We set:
, respectively for observation and background error
variables. We choose here to have uncorrelated errors (that is, diagonal
matrices) and to have the same variance of 1 for all variables (that is,
identity matrices). We set:
B = R = Id = [1 0 0 ; 0 1 0 ; 0 0 1]
Last, we need an observation operator  to convert the
background value in the space of observation values. Here, because the space
dimensions are the same, we can choose the identity as the observation
operator:
to convert the
background value in the space of observation values. Here, because the space
dimensions are the same, we can choose the identity as the observation
operator:
H = Id = [1 0 0 ; 0 1 0 ; 0 0 1]
With such choices, the “Best Linear Unbiased Estimator” (BLUE) will be the
average vector between  and
and  , named the
analysis, denoted by
, named the
analysis, denoted by 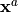 , and its value is:
, and its value is:
Xa = [0.5 0.5 0.5]
As an extension of this example, one can change the variances represented by
 or
or 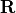 independently, and the analysis
independently, and the analysis
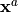 will move to
will move to  or to
or to
 , in inverse proportion of the variances in
, in inverse proportion of the variances in
 and
and 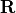 . As an other extension, it is also
equivalent to search for the analysis thought a BLUE algorithm or a 3DVAR one.
. As an other extension, it is also
equivalent to search for the analysis thought a BLUE algorithm or a 3DVAR one.
5.1.2. Using the graphical interface (GUI) to build the ADAO case¶
First, you have to activate the ADAO module by choosing the appropriate module button or menu of SALOME, and you will see:
Choose the “New” button in this window. You will directly get the embedded
case editor interface for variables definition, along with the SALOME “Object
browser”. You can then click on the “New” button  to create a new
ADAO case, and you will see:
to create a new
ADAO case, and you will see:
Then, fill in the variables to build the ADAO case by using the experimental set up described above. All the technical information given above will be directly inserted in the ADAO case definition, by using the String type for each variable. When the case definition is ready, save it to a “JDC (*.comm)” native file somewhere in your path. Remember that other files will be also created near this first one, so it is better to make a specific directory for your case, and to save the file inside. The name of the file will appear in the “Object browser” window, under the “ADAO” menu. The final case definition looks like this:
To go further, we need now to generate the YACS scheme from the ADAO case
definition. In order to do that, right click on the name of the file case in the
“Object browser” window, and choose the “Export to YACS” sub-menu (or the
“Export to YACS” button  ) as below:
) as below:
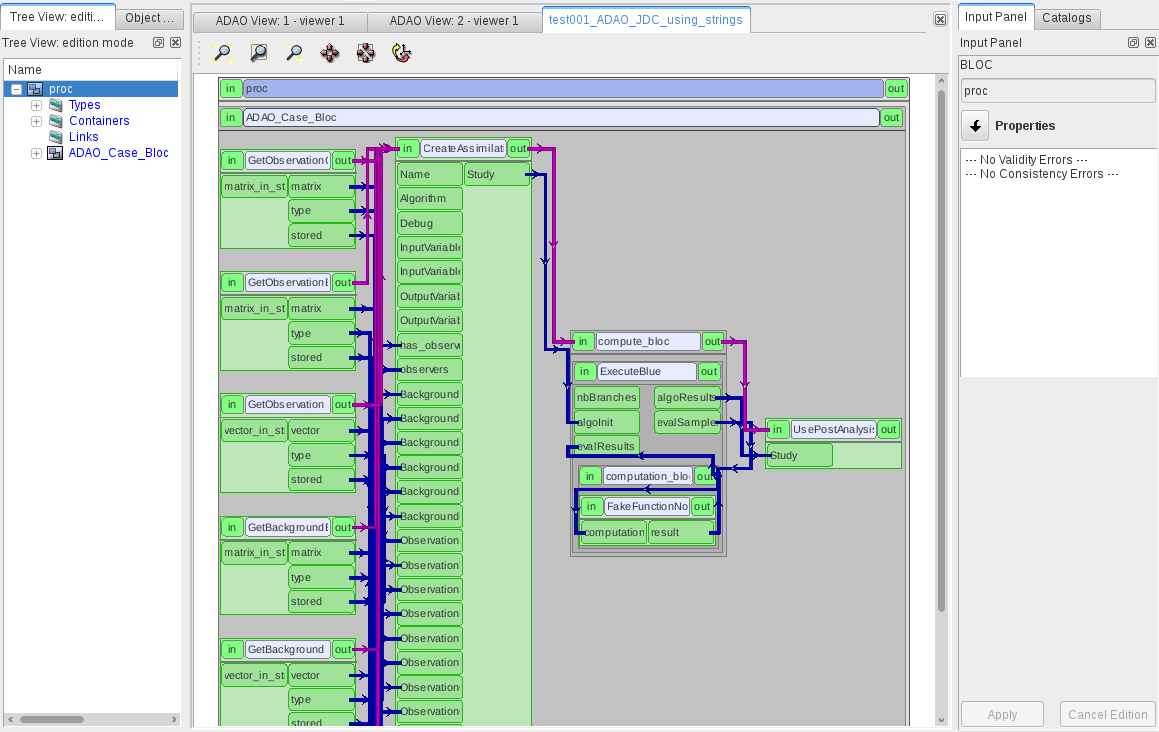
This command will generate the YACS scheme, activate YACS module in SALOME, and open the new scheme in the GUI of the YACS module 1. After eventually reordering the nodes by using the “arrange local nodes” sub-menu of the YACS graphical view of the scheme, you get the following representation of the generated ADAO scheme:
After that point, all the modifications, executions and post-processing of the data assimilation scheme will be done in the YACS module. In order to check the result in a simple way, we can use the “UserPostAnalysis” node (or we create a new YACS node by using the “in-line script node” sub-menu of the YACS graphical view).
This script node will retrieve the data assimilation analysis from the “algoResults” output port of the computation bloc (which gives access to a SALOME Python Object), and will print it on the standard output.
To obtain this, the in-line script node need to have an input port of type “pyobj”, named “Study” for example, that have to be linked graphically to the “algoResults” output port of the computation bloc. Then, the code to fill in the script node is:
Xa = Study.getResults().get("Analysis")[-1]
print()
print("Analysis =",Xa)
print()
The (initial or augmented) YACS scheme can be saved (overwriting the generated scheme if the “Save” command or button are used, or with a new name through the “Save as” command). Ideally, the implementation of such post-processing procedure can be done in YACS to test, and then entirely saved in one Python script that can be integrated in the ADAO case by using the keyword “UserPostAnalysis”.
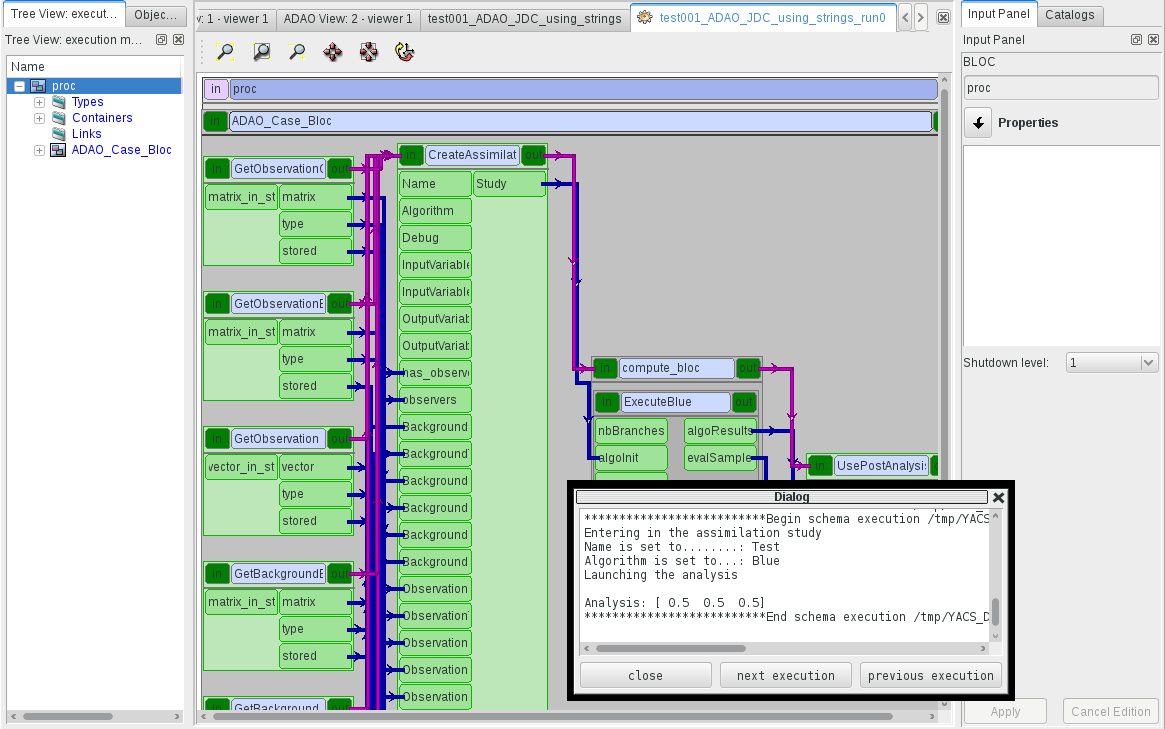
Then, classically in YACS, the scheme have to be compiled for run, and then executed. After completion, the printing on standard output is available in the “YACS Container Log”, obtained through the right click menu of the “proc” window in the YACS scheme as shown below:
We verify that the result is correct by checking that the log dialog window contains information similar to the following line:
Analysis = [0.5, 0.5, 0.5]
as shown in the image above.
As a simple extension of this example, one can notice that the same problem solved with a 3DVAR algorithm gives the same result. This algorithm can be chosen at the ADAO case building step, before entering in YACS step. The ADAO 3DVAR case will look completely similar to the BLUE algorithmic case, as shown by the following figure:
There is only one command changing, with “3DVAR” value in the “Algorithm” field instead of “Blue”.
5.2. Building an estimation case with external data definition by scripts¶
It is useful to get parts or all of the ADAO case data from external definition, using Python script files to provide access to the data. As an example, we build here an ADAO case representing the same experimental setup as in the above example Building an estimation case with explicit data definition, but using data from a single one external Python script file.
First, we write the following script file, using conventional names for the required variables. Here, all the input variables are defined in the same script, but the user can choose to split the file in several ones, or to mix explicit data definition in the ADAO GUI and implicit data definition by external files. The present script file looks like:
import numpy
#
# Definition of the Background as a vector
# ----------------------------------------
Background = [0, 0, 0]
#
# Definition of the Observation as a vector
# -----------------------------------------
Observation = "1 1 1"
#
# Definition of the Background Error covariance as a matrix
# ---------------------------------------------------------
BackgroundError = numpy.array([[1., 0., 0.], [0., 1., 0.], [0., 0., 1.]])
#
# Definition of the Observation Error covariance as a matrix
# ----------------------------------------------------------
ObservationError = numpy.matrix("1 0 0 ; 0 1 0 ; 0 0 1")
#
# Definition of the Observation Operator as a matrix
# --------------------------------------------------
ObservationOperator = numpy.identity(3)
The names of the Python variables above are mandatory, in order to define the right ADAO case variables, but the Python script can be bigger and define classes, functions, file or database access, etc. with other names. Moreover, the above script shows different ways to define arrays and matrices, using list, string (as in Numpy or Octave), Numpy array type or Numpy matrix type, and Numpy special functions. All of these syntax are valid.
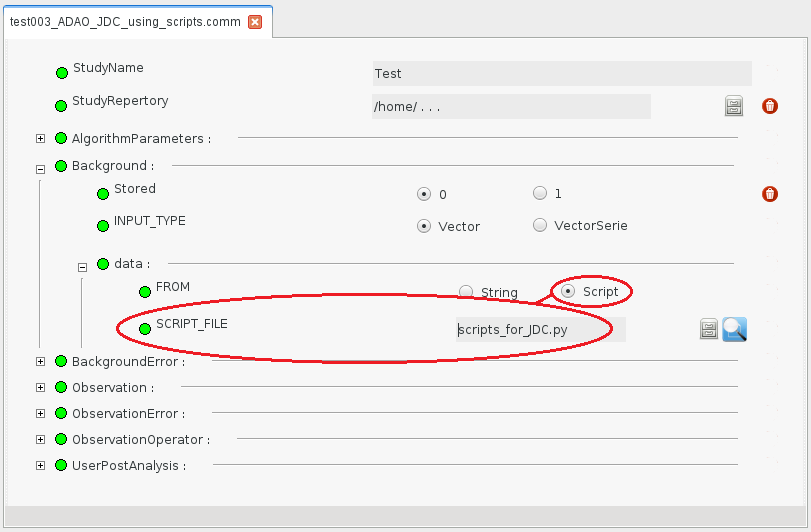
After saving this script in a file (named here “script.py” for the example) somewhere in your path, we use the graphical interface (GUI) to build the ADAO case. The procedure to fill in the case is similar to the previous example except that, instead of selecting the “String” option for the “FROM” keyword of each variable, we select the “Script” one. This leads to a “SCRIPT_DATA/SCRIPT_FILE” entry in the graphical tree, allowing to choose a file as:
Other steps and results are exactly the same as in the Building an estimation case with explicit data definition previous example.
In fact, this script methodology is the easiest way to retrieve data from in-line or previous calculations, from static files, from database or from stream, all of them inside or outside of SALOME. It allows also to modify easily some input data, for example for debug purpose or for repetitive execution process, and it is the most versatile method in order to parametrize the input data. But be careful, script methodology is not a “safe” procedure, in the sense that erroneous data, or errors in calculations, can be directly injected into the ADAO case execution. The user have to carefully verify the content of his scripts.
5.3. Adding parameters to control the data assimilation algorithm¶
One can add some optional parameters to control the data assimilation algorithm calculation. This is done by using optional parameters in the “AlgorithmParameters” command of the ADAO case definition, which is a keyword of the general case command (to choose between “ASSIMILATION_STUDY”, “OPTIMIZATION_STUDY” or “REDUCTION_STUDY”). This keyword requires an explicit definition of the values from default ones, or from a Python dictionary, containing some key/value pairs. The list of possible optional parameters are given in the section [DocR] Reference description of the ADAO commands and keywords and its subsections. The recommendation is to use the explicit definition of values from the default list of optional parameters, as here with the “MaximumNumberOfIterations”:
This dictionary can be defined, for example, in an external Python script file, using the mandatory variable name “AlgorithmParameters” for the dictionary. All the keys inside the dictionary are optional, they all have default values, and can exist without being used. For example:
AlgorithmParameters = {
"Minimizer" : "LBFGSB", # Recommended
"MaximumNumberOfIterations" : 10,
}
If no bounds at all are required on the control variables, then one can choose the “BFGS” or “CG” minimization algorithm for all the variational data assimilation or optimization algorithms. For constrained optimization, the minimizer “LBFGSB” is often more robust, but the “TNC” is sometimes more effective. In a general way, the “LBFGSB” algorithm choice is recommended. Then the script can be added to the ADAO case, in a file entry describing the “Parameters” keyword.
Other steps and results are exactly the same as in the Building an estimation case with explicit data definition previous example. The dictionary can also be directly given in the input field of string type associated for the keyword.
5.4. Building a complex case with external data definition by scripts¶
This more complex and complete example has to been considered as a framework for user inputs treatment, that need to be tailored for each real application. Nevertheless, the file skeletons are sufficiently general to have been used for various applications in neutronic, fluid mechanics… Here, we will not focus on the results, but more on the user control of inputs and outputs in an ADAO case. As previously, all the numerical values of this example are arbitrary.
The objective is to setup the input and output definitions of a physical estimation case by external python scripts, using a general non-linear operator, adding control on parameters and so on… The complete framework scripts can be found in the ADAO skeletons examples directory under the name “External_data_definition_by_scripts”.
5.4.1. Experimental setup¶
We continue to operate in a 3-dimensional space, in order to restrict the size of numerical object shown in the scripts, but the problem is not dependent of the dimension.
We choose a twin experiment context (see the approach
To test a data assimilation chain: the twin experiments), using a known true state  but of arbitrary value:
but of arbitrary value:
Xt = [1 2 3]
The background state  , which represent some a priori
knowledge of the true state, is build as a normal random perturbation of 20% of
the true state
, which represent some a priori
knowledge of the true state, is build as a normal random perturbation of 20% of
the true state  for each component, which is:
for each component, which is:
Xb = Xt + normal(0, 20%*Xt)
To describe the background error covariances matrix  , we make
as previously the hypothesis of uncorrelated errors (that is, a diagonal matrix,
of size 3x3 because
, we make
as previously the hypothesis of uncorrelated errors (that is, a diagonal matrix,
of size 3x3 because  is of length 3) and to have the same
variance of 0.1 for all variables. We get:
is of length 3) and to have the same
variance of 0.1 for all variables. We get:
B = 0.1 * diagonal( length(Xb) )
We suppose that there exist an observation operator  , which
can be non linear. In real calibration procedure or inverse problems, the
physical simulation codes are embedded in the observation operator. We need also
to know its gradient with respect to each calibrated variable, which is a rarely
known information with industrial codes. But we will see later how to obtain an
approximated gradient in this case.
, which
can be non linear. In real calibration procedure or inverse problems, the
physical simulation codes are embedded in the observation operator. We need also
to know its gradient with respect to each calibrated variable, which is a rarely
known information with industrial codes. But we will see later how to obtain an
approximated gradient in this case.
Being in twin experiments, the observation  and its error
covariances matrix
and its error
covariances matrix 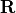 are generated by using the true state
are generated by using the true state
 and the observation operator
and the observation operator  :
:
Yo = H( Xt )
and, with an arbitrary standard deviation of 1% on each error component:
R = 0.0001 * diagonal( length(Yo) )
All the information required for estimation by data assimilation are then defined.
5.4.2. Skeletons of the scripts describing the setup¶
We give here the essential parts of each script used afterward to build the ADAO case. Remember that using these scripts in real Python files requires to correctly define the path to imported modules or codes (even if the module is in the same directory that the importing Python file. One have to mention the encoding if necessary, etc. The indicated file names for the following scripts are arbitrary. Examples of complete file scripts are available in the ADAO examples standard directory.
We first define the true state  and some convenient matrix
building function, in a Python script file named
and some convenient matrix
building function, in a Python script file named
Physical_data_and_covariance_matrices.py:
import numpy
#
def True_state():
"""
Arbitrary values and names, as a tuple of two series of same length
"""
return (numpy.array([1, 2, 3]), ['Para1', 'Para2', 'Para3'])
#
def Simple_Matrix( size, diagonal=None ):
"""
Diagonal matrix, with either 1 or a given vector on the diagonal
"""
if diagonal is not None:
S = numpy.diagflat( diagonal )
else:
S = numpy.identity(int(size))
return S
We can then define the background state  as a random
perturbation of the true state, adding a required ADAO variable at the end of
the script the definition, in order to export the defined value. It is done in a
Python script file named
as a random
perturbation of the true state, adding a required ADAO variable at the end of
the script the definition, in order to export the defined value. It is done in a
Python script file named Script_Background_xb.py:
from Physical_data_and_covariance_matrices import True_state
import numpy
#
xt, names = True_state()
#
Standard_deviation = 0.2*xt # 20% for each variable
#
xb = xt + abs(numpy.random.normal(0.,Standard_deviation,size=(len(xt),)))
#
# Creating the required ADAO variable
# -----------------------------------
Background = list(xb)
In the same way, we define the background error covariance matrix
 as a diagonal matrix, of the same diagonal length as the
background of the true state, using the convenient function already defined. It
is done in a Python script file named
as a diagonal matrix, of the same diagonal length as the
background of the true state, using the convenient function already defined. It
is done in a Python script file named Script_BackgroundError_B.py:
from Physical_data_and_covariance_matrices import True_state, Simple_Matrix
#
xt, names = True_state()
#
B = 0.1 * Simple_Matrix( size = len(xt) )
#
# Creating the required ADAO variable
# -----------------------------------
BackgroundError = B
To continue, we need the observation operator  as a function
of the state. It is here defined in an external file named
as a function
of the state. It is here defined in an external file named
"Physical_simulation_functions.py", which should contain one function
conveniently named here "DirectOperator". This function is user one,
representing as programming function the  operator. We suppose
this function is then given by the user. A simple skeleton is given here for
convenience:
operator. We suppose
this function is then given by the user. A simple skeleton is given here for
convenience:
def DirectOperator( XX ):
import numpy
""" Direct non-linear simulation operator """
#
# --------------------------------------> EXAMPLE TO BE REMOVED
HX = 1. * numpy.ravel( XX ) # EXAMPLE TO BE REMOVED
# --------------------------------------> EXAMPLE TO BE REMOVED
#
return HX
We does not need the linear companion operators "TangentOperator" and
"AdjointOperator" because they will be approximated using ADAO
capabilities. Detailed information on these operators can be found in the
Requirements for functions describing an operator.
We insist on the fact that these non-linear operator "DirectOperator",
tangent operator "TangentOperator" and adjoint operator
"AdjointOperator" come from the physical knowledge, include the reference
physical simulation code, and have to be carefully setup by the data
assimilation or optimization user. The simulation errors or missuses of the
operators can not be detected or corrected by the data assimilation and
optimization ADAO framework alone.
In this twin experiments framework, the observation  and its
error covariances matrix
and its
error covariances matrix 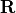 can be generated. It is done in two
Python script files, the first one being named
can be generated. It is done in two
Python script files, the first one being named Script_Observation_yo.py:
from Physical_data_and_covariance_matrices import True_state
from Physical_simulation_functions import DirectOperator
#
xt, noms = True_state()
#
yo = DirectOperator( xt )
#
# Creating the required ADAO variable
# -----------------------------------
Observation = list(yo)
and the second one named Script_ObservationError_R.py:
from Physical_data_and_covariance_matrices import True_state, Simple_Matrix
from Physical_simulation_functions import DirectOperator
#
xt, names = True_state()
#
yo = DirectOperator( xt )
#
R = 0.0001 * Simple_Matrix( size = len(yo) )
#
# Creating the required ADAO variable
# -----------------------------------
ObservationError = R
As in previous examples, it can be useful to define some parameters for the data
assimilation algorithm. For example, if we use the standard “3DVAR” algorithm,
the following parameters can be defined in a Python script file named
Script_AlgorithmParameters.py:
# Creating the required ADAO variable
# -----------------------------------
AlgorithmParameters = {
"Minimizer" : "LBFGSB", # Recommended
"MaximumNumberOfIterations" : 15, # Number of global iterative steps
"Bounds" : [
[ None, None ], # Bound on the first parameter
[ 0., 4. ], # Bound on the second parameter
[ 0., None ], # Bound on the third parameter
],
"StoreSupplementaryCalculations" : [
"CurrentState",
"CostFunctionJ",
],
}
Finally, it is common to post-process the results, retrieving them after the
data assimilation phase in order to analyze, print or show them. It requires to
use a intermediary Python script file in order to extract these results at the
end of the a data assimilation or optimization process. The following example
Python script file, named Script_UserPostAnalysis.py, illustrates the fact:
from Physical_data_and_covariance_matrices import True_state
import numpy
#
xt, names = True_state()
xa = ADD.get("Analysis")[-1]
x_series = ADD.get("CurrentState")[:]
J = ADD.get("CostFunctionJ")[:]
#
# Verifying the results by printing
# ---------------------------------
print()
print("xt = %s"%xt)
print("xa = %s"%numpy.array(xa))
print()
for i in range( len(x_series) ):
print("Step %2i : J = %.5e and X = %s"%(i, J[i], x_series[i]))
print()
At the end, we get a description of the whole case setup through a set of files listed here:
Physical_data_and_covariance_matrices.pyPhysical_simulation_functions.pyScript_AlgorithmParameters.pyScript_BackgroundError_B.pyScript_Background_xb.pyScript_ObservationError_R.pyScript_Observation_yo.pyScript_UserPostAnalysis.py
We insist here that all these scripts are written by the user and can not be automatically tested by ADAO. So the user is required to verify the scripts (and in particular their input/output) in order to limit the difficulty of debug. We recall: script methodology is not a “safe” procedure, in the sense that erroneous data, or errors in calculations, can be directly injected into the YACS scheme execution.
5.4.3. Building the case with external data definition by scripts¶
All these scripts can then be used to define the ADAO case with external data definition by Python script files. It is entirely similar to the method described in the Building an estimation case with external data definition by scripts previous section. For each variable to be defined, we select the “Script” option of the “FROM” keyword, which leads to a “SCRIPT_DATA/SCRIPT_FILE” entry in the tree. For the “ObservationOperator” keyword, we choose the “ScriptWithOneFunction” form and keep the default differential increment.
The other steps to build the ADAO case are exactly the same as in the Building an estimation case with explicit data definition previous section.
Using the simple linear operator  from the Python script file
from the Python script file
Physical_simulation_functions.py in the ADAO examples standard directory,
the results will look like (it may depend on the system):
xt = [1 2 3]
xa = [ 1.000014 2.000458 3.000390]
Step 0 : J = 1.81750e+03 and X = [1.014011, 2.459175, 3.390462]
Step 1 : J = 1.81750e+03 and X = [1.014011, 2.459175, 3.390462]
Step 2 : J = 1.79734e+01 and X = [1.010771, 2.040342, 2.961378]
Step 3 : J = 1.79734e+01 and X = [1.010771, 2.040342, 2.961378]
Step 4 : J = 1.81909e+00 and X = [1.000826, 2.000352, 3.000487]
Step 5 : J = 1.81909e+00 and X = [1.000826, 2.000352, 3.000487]
Step 6 : J = 1.81641e+00 and X = [1.000247, 2.000651, 3.000156]
Step 7 : J = 1.81641e+00 and X = [1.000247, 2.000651, 3.000156]
Step 8 : J = 1.81569e+00 and X = [1.000015, 2.000432, 3.000364]
Step 9 : J = 1.81569e+00 and X = [1.000015, 2.000432, 3.000364]
Step 10 : J = 1.81568e+00 and X = [1.000013, 2.000458, 3.000390]
...
The state at the first step is the randomly generated background state
 . During calculation, these printings on standard output are
available in the “YACS Container Log” window, obtained through the right click
menu of the “proc” window in the YACS executed scheme.
. During calculation, these printings on standard output are
available in the “YACS Container Log” window, obtained through the right click
menu of the “proc” window in the YACS executed scheme.
- 1
For more information on YACS, see the YACS module and its integrated help available from the main menu Help of the SALOME platform.